Web App
Roami
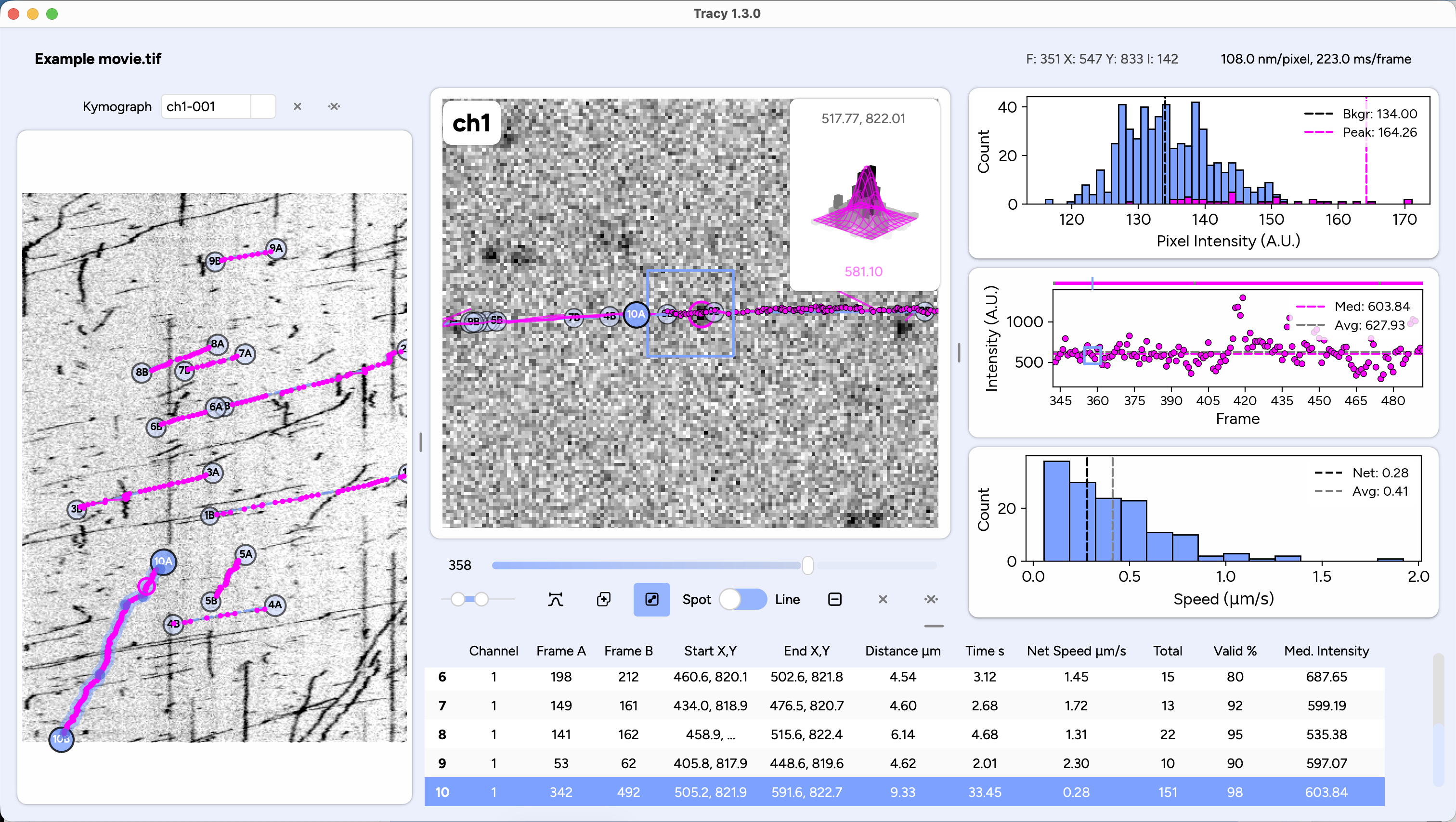
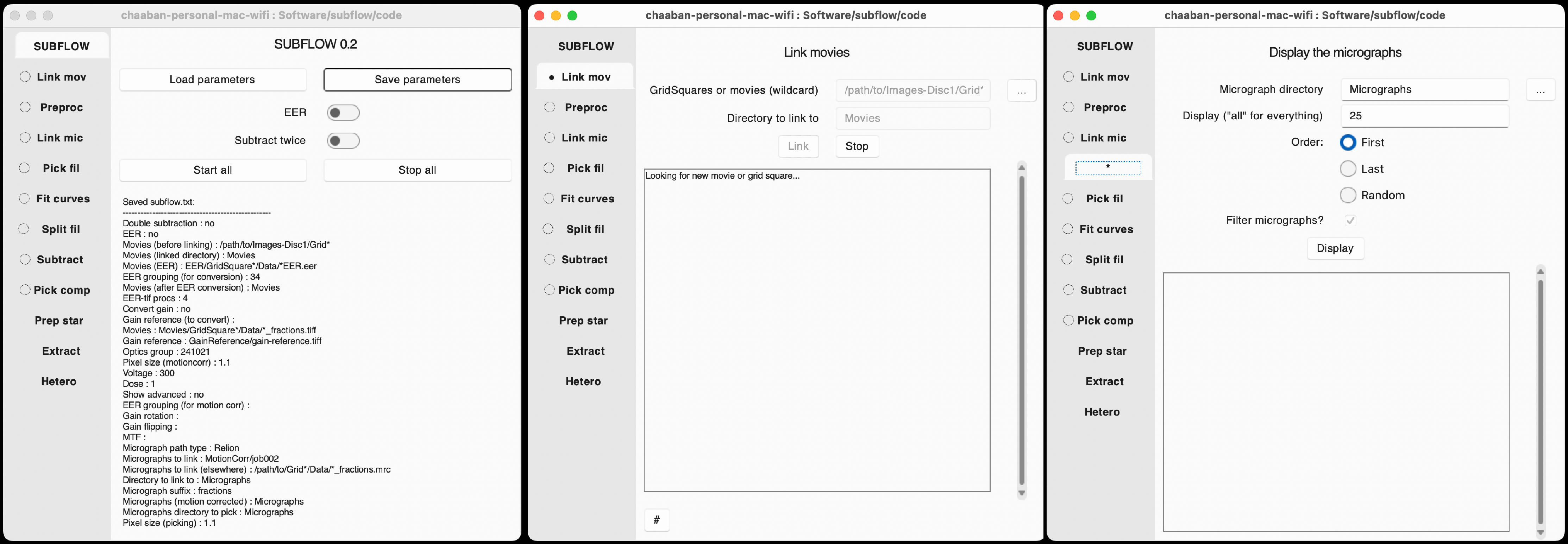
Interactive molecular structure explorer (beta). Analyzes intra- and inter-molecular interactions, can load AlphaFold predictions, browse PAE plots, and includes educational guidance on interaction types, confidence metrics, and amino acid properties. Click here to go to the web app (stripped-down preview on the right).
Built with Three.js, Chapi (Coot), PDBe Arpeggio, Open Babel, and FastAPI.
 GitHub
GitHub
 Download for Mac
Download for Mac
 Download for Windows
Download for Windows